LifeCLEF 2022-23 Plant
Image-based plant identification at global scale
Note: Do not forget to read the Rules section on this page. Pressing the red Participate button leads you to a page where you have to agree with those rules. You will not be able to submit any results before agreeing with the rules.
Note: Before trying to submit results, read the Submission instructions section on this page.
Motivation
It is estimated that there are more than 300,000 species of vascular plants in the world. Increasing our knowledge of these species is of paramount importance for the development of human civilization (agriculture, construction, pharmacopoeia, etc.), especially in the context of the biodiversity crisis. However, the burden of systematic plant identification by human experts strongly penalizes the aggregation of new data and knowledge. Since then, automatic identification has made considerable progress in recent years as highlighted during all previous editions of PlantCLEF. Deep learning techniques now seem mature enough to address the ultimate but realistic problem of global identification of plant biodiversity in spite of many problems that the data may present (a huge number of classes, very strongly unbalanced classes, partially erroneous identifications, duplications, variable visual quality, diversity of visual contents such as photos or herbarium sheets, etc). The PlantCLEF2022&23 challenge editions propose to take a step in this direction by tackling a multi-image (and metadata) classification problem with a very large number of classes (80k plant species).
Data
The training dataset can be distinguished in 2 main categories: "trusted" and "web" (i.e. with or without species labels provided and checked by human experts), totaling 4M images on 80k classes.
The "trusted" training dataset is based on a selection of more than 2.9M images covering 80k plant species shared and collected mainly by GBIF (and EOL to a lesser extent). These images come mainly from academic sources (museums, universities, national institutions) and collaborative platforms such as inaturalist or Pl@ntNet, implying a fairly high certainty of determination quality. Nowadays, many more photographs are available on these platforms for a few thousand species, but the number of images has been globally limited to around 100 images per species, favouring types of views adapted to the identification of plants (close-ups of flowers, fruits, leaves, trunks, ...), in order to not unbalance the classes and to not explode the size of the training dataset.
In contrast, the second data set is based on a collection of web images provided by search engines Google and Bing. This initial collection of several million images suffers however from a significant rate of species identification errors and a massive presence of duplicates and images less adapted for visual identification of plants (herbariums, landscapes, microscopic views...), or even off-topic (portrait photos of botanists, maps, graphs, other kingdoms of the living, manufactured objects, ...). The initial collection has been then semi-automatically revised to drastically reduce the number of these irrelevant pictures and to maximise, as for the trusted dataset, close-ups of flowers, fruits, leaves, trunks, etc. The "web" dataset finally contains about 1.1 million images covering around 57k species..
Lastly, the test set will be a set of tens of thousands pictures verified by world class experts related to various regions of the world and taxonomic groups.
Challenge description
The task will be evaluated as a plant species retrieval task based on multi-image plant observations from the test set. The goal will be to retrieve the correct plant species among the top results of a ranked list of species returned by the evaluated system. The participants will first have access to the training set and a few months later, they will be provided with the whole test set.
Submission instructions
More practically, the run file to be submitted is a csv file (with semicolon separators) and has to contain as much lines as the number of predictions, each prediction being composed of an obsid (the identifier of a plant observation that can be itself composed of several images), a classid, a probability and a rank. Each line should have the following format: <obsid;classid;probability;rank>
Here is a run example respecting this format with random predictions: fake_run
Due to the large number of plant observations in the test set, in order to limit the size of the run files, we ask the participants to limit the predictions to the 30 best results per observation (a maximum of 30 rows from rank 1 to 31 max for each plant observation).
Participants will be allowed to submit a maximum of 10 run files.
As soon as the submission is open, you will find a “Create Submission” button on this page (next to the tabs).
Before being allowed to submit your results, you have to first press the red participate button, which leads you to a page where you have to accept the challenge's rules.
Evaluation criteria:
The primary metrics used for the evaluation of the task will be the Mean Reciprocal Rank. The MRR is a statistic measure for evaluating any process that produces a list of possible responses to a sample of queries ordered by probability of correctness. The reciprocal rank of a query response is the multiplicative inverse of the rank of the first correct answer. The MRR is the average of the reciprocal ranks for the whole test set:

where |Q| is the total number of query occurrences in the test set. However, given the long tail of the data distribution, in order to compensate for species that would be underrepresented in the test set, we will use a macro-averaged version of the MRR (average MRR per species).
Rules
LifeCLEF lab is part of the Conference and Labs of the Evaluation Forum: CLEF. CLEF consists of independent peer-reviewed workshops on a broad range of challenges in the fields of multilingual and multimodal information access evaluation, and a set of benchmarking activities carried in various labs designed to test different aspects of mono and cross-language Information retrieval systems. More details about the conference can be found here.
Submitting a working note with the full description of the methods used in each run is mandatory. Any run that could not be reproduced thanks to its description in the working notes might be removed from the official publication of the results. Working notes are published within CEUR-WS proceedings, resulting in an assignment of an individual DOI (URN) and an indexing by many bibliography systems including DBLP. According to the CEUR-WS policies, a light review of the working notes will be conducted by LifeCLEF organizing committee to ensure quality. As an illustration,LifeCLEF 2021 working notes (task overviews and participant working notes) can be found within CLEF 2021 CEUR-WS proceedings.
Important
Participants of this challenge will automatically be registered at CLEF. In order to be compliant with the CLEF registration requirements, please edit your profile by providing the following additional information:
-
First name
-
Last name
-
Affiliation
-
Address
-
City
-
Country
-
Regarding the username, please choose a name that represents your team.
This information will not be publicly visible and will be exclusively used to contact you and to send the registration data to CLEF, which is the main organizer of all CLEF labs
CEUR Working Notes
For detailed instructions, please refer to https://clef2023.clef-initiative.eu/index.php?page=Pages/publications.html
A summary of the most important points:
- All participating teams with at least one graded submission, regardless of the score, should submit a CEUR working notes paper.
- Submission of reports is done through EasyChair – please make absolutely sure that the author (names and order), title, and affiliation information you provide in EasyChair match the submitted PDF exactly
- Strict deadline for Working Notes Papers: 7 June 2023
- Strict deadline for CEUR-WS Camera Ready Working Notes Papers: 7 July 2023
- Templates are available here
- Working Notes Papers should cite both the LifeCLEF 2023 overview paper as well as the PlantCLEF task overview paper, citation information will be added in the Citations section below as soon as the titles have been finalized.
Schedule
- Jan 2023: registration opens for all LifeCLEF challenges
- 1 February 2023: training data release
- 15 March 2023: test data release and opening of the submission system
- 22 May 2023: closing the submission system
- 26 May 2023: release of processed results by the task organizers
- 7 June 2023: deadline for submission of working notes papers by the participants
- 23 June 2023: notification of acceptance of working note papers [CEUR-WS proceedings]
- 7 July 2023: camera ready working notes papers of participants and organizers
- 18-21 Sept 2023: CLEF 2023 Thessaloniki
Participation Requirements
Participants must register to LifeCLEF – PlantCLEF2023 task by filling out this form:
https://clef2023-labs-registration.dei.unipd.it/registrationForm.php
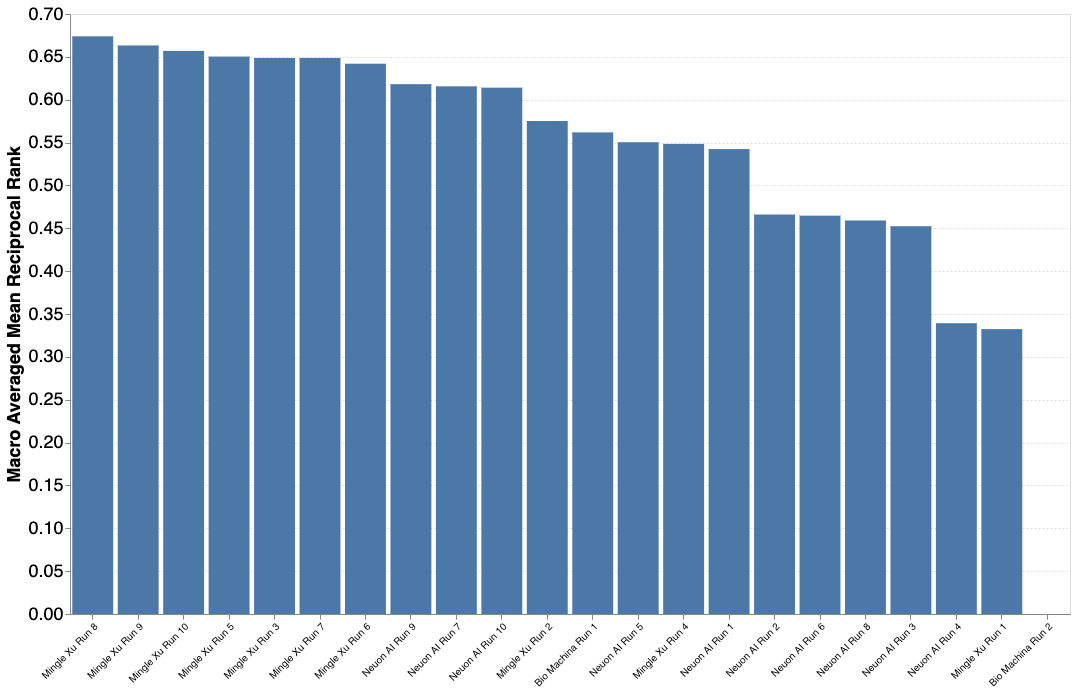
Results PlantCLEF 2023
A total of 3 participants submitted 22 runs. The results are encouraging despite the great difficulty of the challenge! Thanks again for all your efforts and your investment on this problem of great importance for a better knowledge of the biodiversity of plants.
| Team run name | Aicrowd name | Filename | MA-MRR |
|---|---|---|---|
| Mingle Xu Run 8 | MingleXu | eva_l_psz14_21k_ft_psz14to16_TrainTrustWeb_epoch99_sorted_top30 | 0.67395 |
| Mingle Xu Run 9 | MingleXu | eva_l_psz14_21k_ft_psz14to16_TrainTrustWeb_epoch99_FT_Trust_epoch9_sorted_top30 | 0.66330 |
| Mingle Xu Run 10 | MingleXu | eva_l_psz14_21k_ft_psz14to16_TrainTrustWeb_epoch99_FT_Trust_epoch49_sorted_top30 | 0.65695 |
| Mingle Xu Run 5 | MingleXu | eva_l_psz14to16_21k_ft_epoch99_sorted_top30 | 0.65035 |
| Mingle Xu Run 3 | MingleXu | eva_l_psz14to16_epoch99_sorted_top30 | 0.64871 |
| Mingle Xu Run 7 | MingleXu | eva_l_psz14_21k_ft_psz14to16_TrainTrustWeb_epoch99_sorted_top30 | 0.64871 |
| Mingle Xu Run 6 | MingleXu | eva_l_psz14_21k_ft_psz14to16_stat7_54478cls_2807969img_epoch99_sorted_top30 | 0.64201 |
| Neuon AI Run 9 | neuon_ai | 9_run9_both_random_ens_2022 | 0.61813 |
| Neuon AI Run 7 | neuon_ai | 7_run9_cont_random_ens_2022 | 0.61561 |
| Neuon AI Run 10 | neuon_ai | 10_run9_cont_random_ens_2022 | 0.61406 |
| Mingle Xu Run 2 | MingleXu | eva_l_psz14_21k_ft_psz14to16_stat36_28681cls_2381264img_epoch99_sorted_top30 | 0.57514 |
| Bio Machina Run 1 | BioMachina | vit_base_patch16_224-1ouumtje-epoch=15-train_loss=0.23-train_acc=0.82--val_loss=0.56-val_acc=0.78.ckpt | 0.56186 |
| Neuon AI Run 5 | neuon_ai | 5_run9_cont_random | 0.55040 |
| Mingle Xu Run 4 | MingleXu | eva_l_psz14_21k_ft_psz14to16_stat50_24284cls_2194912img_epoch99_sorted_top30 | 0.54846 |
| Neuon AI Run 1 | neuon_ai | 1_run9_random | 0.54242 |
| Neuon AI Run 2 | neuon_ai | 2_run5_predefined | 0.46606 |
| Neuon AI Run 6 | neuon_ai | 6_run9_cont_random_featurematch | 0.46476 |
| Neuon AI Run 8 | neuon_ai | 8_run9_cont_random_featurematch_w | 0.45910 |
| Neuon AI Run 3 | neuon_ai | 3_run9_random_featurematch | 0.45242 |
| Neuon AI Run 4 | neuon_ai | 4_organs_model | 0.33926 |
| Mingle Xu Run 1 | MingleXu | eva_l_psz14_21k_ft_psz14to16_stat100_9122cls_920774img_epoch99_sorted_top30 | 0.33239 |
| Bio Machina Run 2 | BioMachina | vit_base_patch16_224-hkgh1i0s-epoch=19-train_loss=0.09-train_acc=0.92--val_loss=0.43-val_acc=0.87.ckpt | 0.00000 |
------------------------------------------------------------------------------------------------------
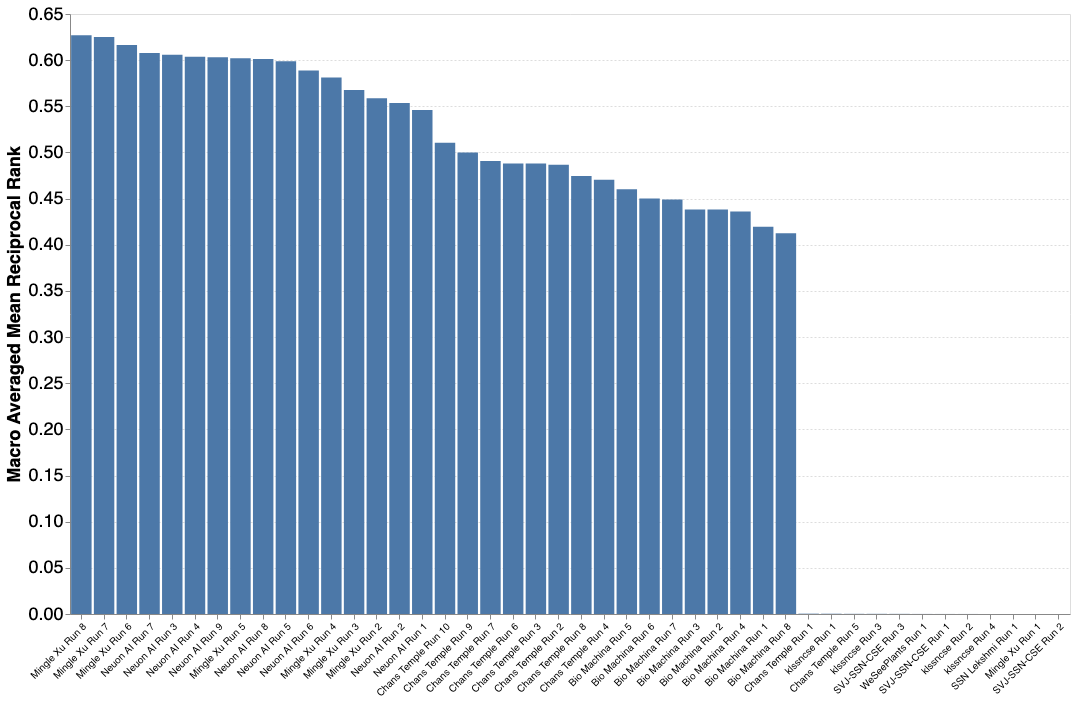
Results PlantCLEF 2022 (Previous round)
A total of 8 participants submitted 45 runs. The results are encouraging despite the great difficulty of the challenge! Thanks again for all your efforts and your investment on this problem of great importance for a better knowledge of the biodiversity of plants.
| Team run name | Aicrowd name | Filename | MA-MRR |
|---|---|---|---|
| Mingle Xu Run 8 | MingleXu | submission_epoch80 | 0.62692 |
| Mingle Xu Run 7 | MingleXu | submission_epoch77 | 0.62497 |
| Mingle Xu Run 6 | MingleXu | submission_epoch67 | 0.61632 |
| Neuon AI Run 7 | neuon_ai | 7_trusted_with_trusted_and_web_inceptionres_inception_ens | 0.60781 |
| Neuon AI Run 3 | neuon_ai | 3_trusted_and_web_inceptionres_inception_ens | 0.60583 |
| Neuon AI Run 4 | neuon_ai | 4_trusted_with_trusted_and_web_inceptionres_inception_ens | 0.60381 |
| Neuon AI Run 9 | neuon_ai | 9_trusted_with_trusted_and_web_inceptionres_inception_ens | 0.60301 |
| Mingle Xu Run 5 | MingleXu | submission_epoch49 | 0.60219 |
| Neuon AI Run 8 | neuon_ai | 8_trusted_with_trusted_and_web_inceptionres_inception_ens_pretained | 0.60113 |
| Neuon AI Run 5 | neuon_ai | 5_trusted_and_web_inceptionres_inception_ens_triplet_dictionary | 0.59892 |
| Neuon AI Run 6 | neuon_ai | 6_trusted_with_trusted_and_web_inceptionres_ens | 0.58874 |
| Mingle Xu Run 4 | MingleXu | submission_epoch24 | 0.58110 |
| Mingle Xu Run 3 | MingleXu | submission_epoch15 | 0.56772 |
| Mingle Xu Run 2 | MingleXu | submission | 0.55865 |
| Neuon AI Run 2 | neuon_ai | 2_trusted_inceptionres_inception_ens | 0.55358 |
| Neuon AI Run 1 | neuon_ai | 1_trusted_inceptionres_5_labels | 0.54613 |
| Chans Temple Run 10 | Chans_Temple_1 | rs34n50e_attempt_lookup_ip_agg | 0.51043 |
| Chans Temple Run 9 | Chans_Temple_1 | rs34e_attempt_lookup_ip_agg | 0.49994 |
| Chans Temple Run 7 | Chans_Temple_1 | naive_attempt_lookup_i_agg | 0.49075 |
| Chans Temple Run 6 | Chans_Temple_1 | naive_attempt_lookup_agg | 0.48804 |
| Chans Temple Run 3 | Chans_Temple_1 | naive_attempt_lookup_agg | 0.48804 |
| Chans Temple Run 2 | Chans_Temple_1 | naive_attempt | 0.48661 |
| Chans Temple Run 8 | Chans_Temple_1 | hydra_attempt_lookup_i_agg | 0.47447 |
| Chans Temple Run 4 | Chans_Temple_1 | naive_attempt_logit_agg | 0.47034 |
| Bio Machina Run 5 | BioMachina | results-resnet50-webpretrained-trusted-epoch25 | 0.46010 |
| Bio Machina Run 6 | BioMachina | resnet101-epoch=7-step=180424-web.trusted | 0.45011 |
| Bio Machina Run 7 | BioMachina | resnet101-epoch=10-step=248083-web-trusted | 0.44910 |
| Bio Machina Run 3 | BioMachina | results-3 | 0.43820 |
| Bio Machina Run 2 | BioMachina | results | 0.43813 |
| Bio Machina Run 4 | BioMachina | results-resnet50-webpretrained-trusted-epoch10 | 0.43606 |
| Bio Machina Run 1 | BioMachina | best_supr_hefficientnet_b4-ds=trusted-epoch=15-train_loss=1.81-train_acc=0.57--val_loss=2.33-val_acc=0.49 | 0.41950 |
| Bio Machina Run 8 | BioMachina | results | 0.41240 |
| Chans Temple Run 1 | Chans_Temple_1 | sanity_baseline | 0.00036 |
| klssncse Run 1 | klssncse | output | 0.00029 |
| Chans Temple Run 5 | Chans_Temple_1 | naive_attempt_lookup_i_agg | 0.00019 |
| klssncse Run 3 | klssncse | submission-kl | 0.00018 |
| SVJ-SSN-CSE Run 3 | SVJ-SSN-CSE | res_final | 0.00015 |
| WeSeePlants Run 1 | WeSeePlants | FinalResult | 0.00007 |
| klssncse Run 2 | klssncse | FinalSubmission_File | 0.00005 |
| SVJ-SSN-CSE Run 1 | SVJ-SSN-CSE | res | 0.00005 |
| klssncse Run 4 | klssncse | submission-kl | 0.00003 |
| SSN Lekshmi Run 1 | SSN_Lekshmi | submission-kl | 0.00003 |
| Mingle Xu Run 1 | MingleXu | results | 0.00003 |
| SVJ-SSN-CSE Run 2 | SVJ-SSN-CSE | res | 0.00000 |
Citations
Information will be posted after the challenge ends.
Prizes
Publication
LifeCLEF 2023 is an evaluation campaign that is being organized as part of the CLEF initiative labs. The campaign offers several research tasks that welcome participation from teams around the world. The results of the campaign appear in the working notes proceedings, published by CEUR Workshop Proceedings (CEUR-WS.org). Selected contributions among the participants, will be invited for publication in the following year in the Springer Lecture Notes in Computer Science (LNCS) together with the annual lab overviews.
Resources
Contact us
Discussion Forum
- You can ask questions related to this challenge on the Discussion Forum. Before asking a new question please make sure that question has not been asked before.
- Click on Discussion tab above or direct link: https://www.aicrowd.com/challenges/lifeclef-2022-23-plant/discussion
Alternative channels
We strongly encourage you to use the public channels mentioned above for communications between the participants and the organizers. In extreme cases, if there are any queries or comments that you would like to make using a private communication channel, then you can send us an email at :
- herve [dot] goeau [at] cirad [dot] fr
- pierre [dot] bonnet [at] cirad [dot] fr
- alexis [dot] joly [at] inria [dot] fr
More information
You can find additional information on the challenge here: https://www.aicrowd.com/challenges/lifeclef-2022-23-plant/discussion
Participants



















































 Continue with Google
Continue with Google
 Sign Up with Email
Sign Up with Email